Introduction
While the surrogate modelling approaches such as Kriging and Polynomial Chaos Expansion can handle problems with low to medium dimensional random input parameters, their accuracy is affected when the number of input dimensions/parameters/features is high- "Curse of dimensionality". To make the UQ and optimization based on high dimensional data with a large number of features, it is paramount to identify the influential features in the hidden/latent space, and perform UQ and optimization based on the latent space.
One of the widely used approaches for dimensionality reduction is principal component analysis (PCA). However, the PCA approach is a linear transformation from the high dimensional data along the dimensions that captures the maximum variance, and is inaccurate when the embedded/latent space is infact non-linear. To tackle this, various manifold learning approaches such as locally-linear embedding, t-stochastic neighbor embedding (t-SNE), and most recently uniform manifold approximation and projection (UMAP) are available, which are also known as non-linear dimensionality reduction techniques.
Currently, I am using these manifold learning and autoencoders based dimensionality reduction for uncertainty quantification and global sensitivity analysis of multi-response system.
One of the widely used approaches for dimensionality reduction is principal component analysis (PCA). However, the PCA approach is a linear transformation from the high dimensional data along the dimensions that captures the maximum variance, and is inaccurate when the embedded/latent space is infact non-linear. To tackle this, various manifold learning approaches such as locally-linear embedding, t-stochastic neighbor embedding (t-SNE), and most recently uniform manifold approximation and projection (UMAP) are available, which are also known as non-linear dimensionality reduction techniques.
Currently, I am using these manifold learning and autoencoders based dimensionality reduction for uncertainty quantification and global sensitivity analysis of multi-response system.
|
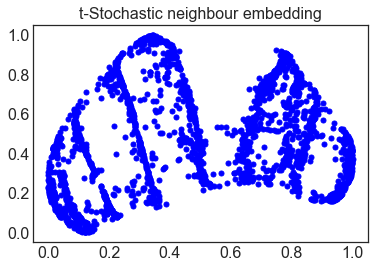
This widely used benchmark problem demonstrates the dimensionality reduction using principal-component analysis and manifold learning techniques such as LLE, t-SNE, and UMAP.
As seen on the right hand figures, the dimensionality reduction with PCA is inappropriate since the most of the data points in higher-dimensions are found to be overlapping in the lower dimensional latent space. Comparatively, the data points formed a well-defined structure with almost no overlapping of the higher dimensional data points in the latent space. Note: Most of these manifold learning techniques are widely available in python packages such as Sklearn. |
Application: Manifold Learning of Progressive Failure Responses of Composite Structures
 Different realizations of load-displacement curves
Different realizations of load-displacement curves
In this ongoing study, the progressive failure responses of composite structures is considered. During progressive failure analysis (PFA), each of the load steps yields a reaction load. Accounting for a large number of load steps, thus create a multi-response problem, and the response is vector valued. Therefore, the goal is to find the latent space using manifold learning for PFA of composite structures.
As can be seen below, the UMAP is able to create an untangle structure unlike the PCA, LLE, and t-SNE when the number of dimensions is reduced from D=40 to d=2.
Note: Higher number of latent space dimensions may be required to obtain better information.
Using number of reduced dimensions in the latent space, d=2 was able to capture only 67% covariance of the random responses with PCA whereas using d=5 was able to capture 99.9% of the covariance. This method of checking the variance with the reduced dimensions guides us in selecting the number of reduced dimensions.
As can be seen below, the UMAP is able to create an untangle structure unlike the PCA, LLE, and t-SNE when the number of dimensions is reduced from D=40 to d=2.
Note: Higher number of latent space dimensions may be required to obtain better information.
Using number of reduced dimensions in the latent space, d=2 was able to capture only 67% covariance of the random responses with PCA whereas using d=5 was able to capture 99.9% of the covariance. This method of checking the variance with the reduced dimensions guides us in selecting the number of reduced dimensions.